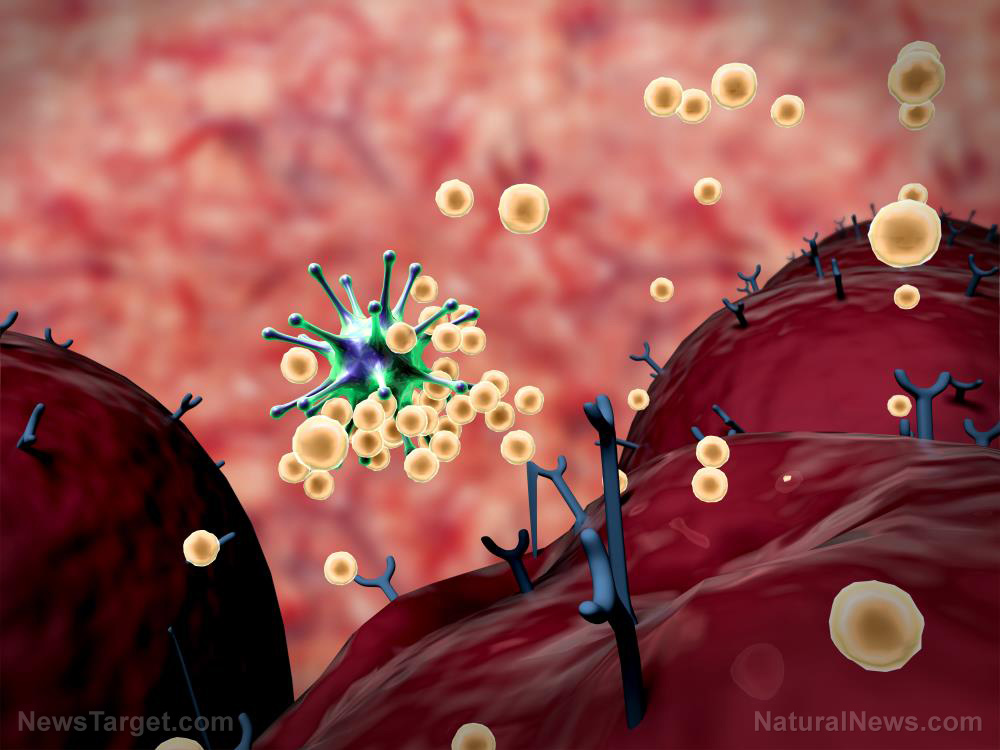
How coronaviruses infect their target cells
The coronavirus family consists of a number of enveloped viruses that contain genetic material known as RNA. Unlike other microscopic pathogens, viruses are not cells; they, therefore, lack the means to proliferate by themselves. To survive and replicate, viruses need to hijack cells and make use of their machinery. They do so by first attaching to the surfaces of target cells and injecting their genetic material into them in a step-by-step process commonly known as infection.
Viral attachment to cells is facilitated by the interaction between proteins found on cell surfaces (receptors) and proteins present on the viral envelope. In coronaviruses, these proteins are known as spike glycoproteins, or simply S protein. The S protein contains two subunits, S1 and S2; while S1 enables coronaviruses to bind to cell surface receptors, S2 makes virus-cell fusion possible.
Two types of cell receptors, namely, angiotensin-converting enzyme 2 (ACE2) and dipeptidyl peptidase 4 (DPP4), recognize a specific part of the S1 subunit known as the receptor-binding domain (RBD). This is particularly true for the S proteins of SARS-CoV-1, the coronavirus behind the 2003 SARS outbreak, and MERS-CoV, the virus that caused the 2015 and 2018 MERS outbreaks. According to the latest studies, SARS-CoV-2, the virus responsible for the current pandemic, specifically binds to ACE2 to gain entry into target cells.
Why antibodies derived from llamas?
Back in 2016, some of the study authors, particularly those from the University of Ghent in Belgium, began isolating antibodies from animals in hopes of developing potential treatments for the diseases caused by SARS-CoV-1 and MERS-CoV. But instead of the usual rabbits or horses, the researchers opted to use llamas due to the unique antibodies they produce in response to infection by microbial pathogens.
In the presence of bacteria or viruses, the immune systems of camelids, such as camels, llamas and alpacas, generate two types of antibodies: one very similar to those produced by humans, and the other, merely a protein fragment (peptide) about 110 amino acids long. This peptide came to be known as single-domain antibody or VHH, because it contains only a single variable domain instead of the usual two.
The VHHs produced by camelids are the smallest naturally derived antibodies that can bind to specific antigens just as efficiently as full-sized ones. VHHs are also very easy to engineer, economical to produce and possess higher thermal and chemical stability than conventional antibodies. Because of these advantageous properties, scientists have been studying VHHs for use against respiratory infections for years.
"The use of VHHs as biologics in the context of a respiratory infection is a particularly attractive application, since the highly stable VHHs can be nebulized and administered via an inhaler directly to the site of infection," the researchers wrote in their report. "Moreover, due to their stability after prolonged storage, VHHs could be stockpiled as therapeutic treatment options in case of an epidemic."
How VHHs can prevent the coronavirus from entering susceptible cells
In their 2016 study, the researchers were able to isolate an antibody they called VHH-72 from a then-9-months old llama named Winter. Winter still lives on a farm in the Belgian countryside together with some 130 other llamas and alpacas. To make her produce antibodies against SARS-CoV-1 and MERS-CoV, the researchers injected her with pseudotyped viruses, which are stabilized forms of the two coronaviruses that can't cause sickness but still produces S proteins.
The researchers found that VHH-72 has a strong affinity to the SARS-CoV-1 S protein, which caused it to bind tightly to the protein and prevent the virus from attaching to its target cells. When the current pandemic erupted, they immediately wondered if VHH-72 could also bind to the S protein of SARS-CoV-2, which shares similarities with its cousin, SARS-CoV-1. Although VHH-72 did bind to the SARS-CoV-2 S protein, its affinity to the protein's RBD was rather weak.
To enhance its affinity, the researchers decided to create two variants of VHH-72. One was a fusion of two VHH-72s, while the other was a copy of VHH-72 fused together with the crystallizable fragment (Fc) of the human IgG1 antibody. The researchers reported that these two variants successfully reduced the binding of SARS-CoV-2 to the ACE2 receptors of cultured cells. They accomplished this by binding to the coronavirus's S protein themselves. The two engineered antibodies did the same thing when tested against SARS-CoV-1, making them the first antibodies to neutralize both coronaviruses.
With the success of this experiment, the researchers are now preparing to conduct preclinical trials in animals. This next step will bring them closer to their goal of testing the antibodies in humans. If the antibodies show the same efficiency in humans, then scientists can use them to develop a treatment that will finally put an end to coronavirus pandemic. An antibody-based treatment will also prove more useful than a coronavirus vaccine. (Related: New research suggests vitamin D may help combat coronavirus.)
"Vaccines have to be given a month or two before infection to provide protection," said Jason McLellan, an associate professor of molecular biosciences at The University of Texas at Austin and co-senior author of the study. "With antibody therapies, you're directly giving somebody the protective antibodies and so, immediately after treatment, they should be protected. The antibodies could also be used to treat somebody who is already sick to lessen the severity of the disease."
Sources include:
Please contact us for more information.























