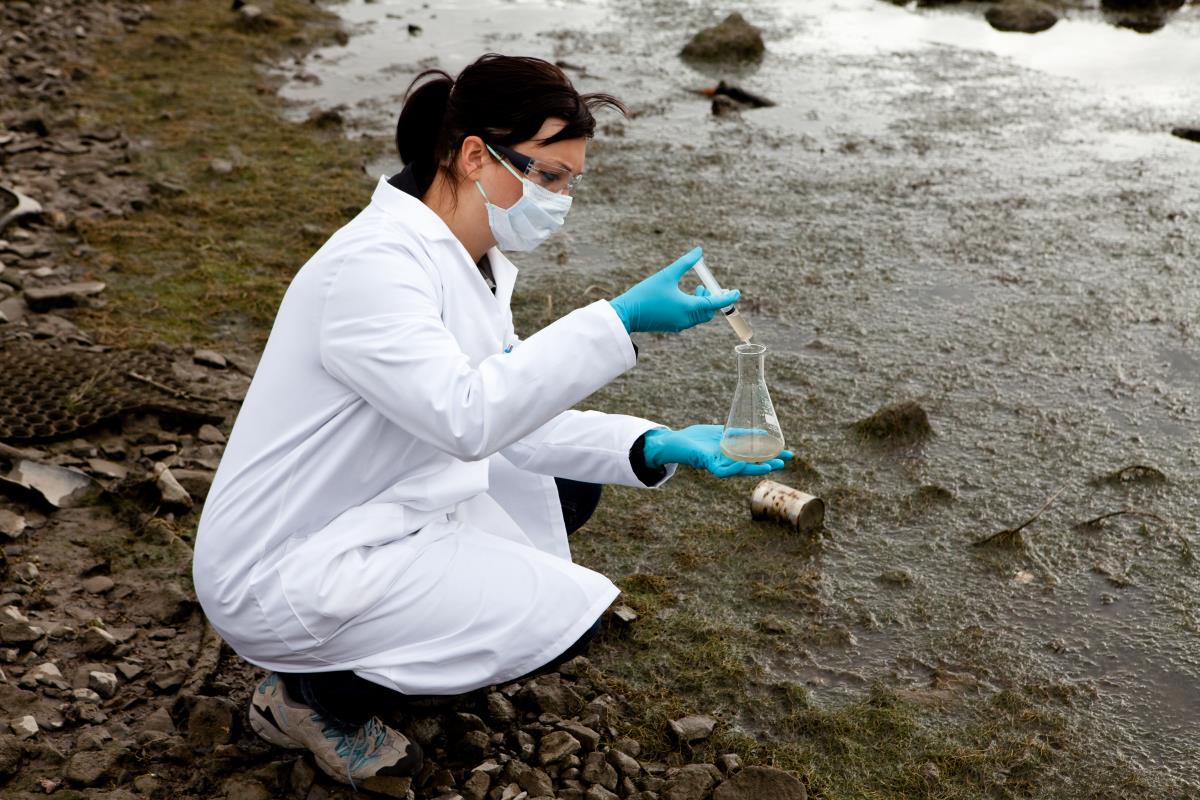
The drawbacks of conventional fecal-source detection methods
Currently used methods for identifying sources of fecal contamination in water are called microbial source tracking (MST) methods. MST methods are used to manage water quality and assess public health risk. MST methods make use of the association of certain fecal microorganisms with a specific host to discriminate between different fecal pollution sources. For instance, bacteria from the order Bacteroidales, which are abundant in mammalian wastes, are often used as fecal indicators.
"Certain bacteria are only found in the faeces of very specific species of animal. When analysing DNA samples of these bacteria, you can pinpoint exactly which creatures are the source of the contamination," explains Georg Reischer, one of the authors of the study.
"There are, for example, bacteria which are commonly found in the intestinal microbiome of ruminants. If this type of DNA is found in a water sample, it is highly likely that the contamination originated from grazing cattle." (Related: Are antibiotic residues and fecal pollution responsible for the increased level of antibiotic-resistant bacteria in the environment?)
To detect and quantify genetic markers from water samples, quantitative polymerase chain reaction (qPCR) is used. PCR is a molecular biology technique that allows researchers to make multiple copies of a certain DNA for various analyses. While this method facilitates the identification of fecal pollution sources, it also requires costly equipment with very high precision, which limits its application. qPCR machines also need people with extensive technical training to run them and interpret the results.
A simple DNA-detection method that works like a pregnancy test
To address the limitations of current MST methods, the researchers developed a detection technique that eliminates the need for heavy equipment, is much simpler to execute, and can quickly produce results. This method combines helicase-dependent amplification (HDA) with a strip test to detect fecal pollution caused by ruminants. Unlike qPCR, which requires temperature changes to amplify DNA strands successfully, HDA only needs one constant temperature to drive the entire process.
For the HDA-strip assay, the researchers used a heating block to make copies of a ruminant-associated marker called Bacteroidetes 16S rRNA marker (BacR). Afterward, they applied the reaction mixture onto a test strip that uses a colorimetric reaction to detect and display the amplified DNA products by marker-specific hybridization probes. The researchers reported that the whole assay only takes two hours to complete and can be performed by anyone without the need for extensive training.
To evaluate its efficacy, the researchers did a trial and compared the results of their assay with that of a qPCR reference. Their BacR HDA-strip assay performed comparably well in terms of source-sensitivity, source-specificity, analytical limit of detection, and sample limit of detection. The researchers were quick to note that, while their new method can only produce qualitative results, it can "pave the way for future simple-to-use-MST screening tools."
"Fundamentally, this technology is applicable to many different bacteria and viruses, but for now we are concentrating on the detection of dangerous microbes in water, this being a particularly widespread problem," said Reischer.
He and his team are currently developing a prototype and searching for a suitable industry partner to collaborate with.
Sources include:
Please contact us for more information.